Combination and Permutation Basics
Joseph Wood
2026-03-04
Source:vignettes/GeneralCombinatorics.Rmd
GeneralCombinatorics.RmdThis document serves as an overview for attacking common
combinatorial problems in R. One of the goals of
RcppAlgos is to provide a comprehensive and accessible
suite of functionality so that users can easily get to the heart of
their problem. As a bonus, the functions in RcppAlgos are
extremely efficient and are constantly being improved with every
release.
It should be noted that this document only covers common problems.
For more information on other combinatorial problems addressed by
RcppAlgos, see the following vignettes:
- Combinatorial Sampling
- Constraints in RcppAlgos: Constraint-Driven Combinatorial Enumeration
- Attacking Problems Related to the Subset Sum Problem
- Combinatorial Iterators in RcppAlgos
For much of the output below, we will be using the following function obtained here combining head and tail methods in R (credit to user @flodel)
ht <- function(d, m = 5, n = m) {
## print the head and tail together
cat("head -->\n")
print(head(d, m))
cat("--------\n")
cat("tail -->\n")
print(tail(d, n))
}Introducing comboGeneral and
permuteGeneral
Easily executed with a very simple interface. The output is in lexicographical order.
We first look at getting results without repetition. You can pass an
integer n and it will be converted to the sequence
1:n, or you can pass any vector with an atomic type
(i.e. logical, integer, numeric,
complex, character, and raw).
library(RcppAlgos)
options(width = 90)
## combn output for reference
combn(4, 3)
#> [,1] [,2] [,3] [,4]
#> [1,] 1 1 1 2
#> [2,] 2 2 3 3
#> [3,] 3 4 4 4
## This is the same as combn expect the output is transposed
comboGeneral(4, 3)
#> [,1] [,2] [,3]
#> [1,] 1 2 3
#> [2,] 1 2 4
#> [3,] 1 3 4
#> [4,] 2 3 4
## Find all 3-tuple permutations without
## repetition of the numbers c(1, 2, 3, 4).
head(permuteGeneral(4, 3))
#> [,1] [,2] [,3]
#> [1,] 1 2 3
#> [2,] 1 2 4
#> [3,] 1 3 2
#> [4,] 1 3 4
#> [5,] 1 4 2
#> [6,] 1 4 3
## If you don't specify m, the length of v (if v is a vector) or v (if v is a
## scalar (see the examples above)) will be used
v <- c(2, 3, 5, 7, 11, 13)
comboGeneral(v)
#> [,1] [,2] [,3] [,4] [,5] [,6]
#> [1,] 2 3 5 7 11 13
head(permuteGeneral(v))
#> [,1] [,2] [,3] [,4] [,5] [,6]
#> [1,] 2 3 5 7 11 13
#> [2,] 2 3 5 7 13 11
#> [3,] 2 3 5 11 7 13
#> [4,] 2 3 5 11 13 7
#> [5,] 2 3 5 13 7 11
#> [6,] 2 3 5 13 11 7
## They are very efficient...
system.time(comboGeneral(25, 12))
#> user system elapsed
#> 0.046 0.007 0.052
comboCount(25, 12)
#> [1] 5200300
ht(comboGeneral(25, 12))
#> head -->
#> [,1] [,2] [,3] [,4] [,5] [,6] [,7] [,8] [,9] [,10] [,11] [,12]
#> [1,] 1 2 3 4 5 6 7 8 9 10 11 12
#> [2,] 1 2 3 4 5 6 7 8 9 10 11 13
#> [3,] 1 2 3 4 5 6 7 8 9 10 11 14
#> [4,] 1 2 3 4 5 6 7 8 9 10 11 15
#> [5,] 1 2 3 4 5 6 7 8 9 10 11 16
#> --------
#> tail -->
#> [,1] [,2] [,3] [,4] [,5] [,6] [,7] [,8] [,9] [,10] [,11] [,12]
#> [5200296,] 13 14 15 16 18 19 20 21 22 23 24 25
#> [5200297,] 13 14 15 17 18 19 20 21 22 23 24 25
#> [5200298,] 13 14 16 17 18 19 20 21 22 23 24 25
#> [5200299,] 13 15 16 17 18 19 20 21 22 23 24 25
#> [5200300,] 14 15 16 17 18 19 20 21 22 23 24 25
## And for permutations... over 8 million instantly
system.time(permuteGeneral(13, 7))
#> user system elapsed
#> 0.018 0.007 0.024
permuteCount(13, 7)
#> [1] 8648640
ht(permuteGeneral(13, 7))
#> head -->
#> [,1] [,2] [,3] [,4] [,5] [,6] [,7]
#> [1,] 1 2 3 4 5 6 7
#> [2,] 1 2 3 4 5 6 8
#> [3,] 1 2 3 4 5 6 9
#> [4,] 1 2 3 4 5 6 10
#> [5,] 1 2 3 4 5 6 11
#> --------
#> tail -->
#> [,1] [,2] [,3] [,4] [,5] [,6] [,7]
#> [8648636,] 13 12 11 10 9 8 3
#> [8648637,] 13 12 11 10 9 8 4
#> [8648638,] 13 12 11 10 9 8 5
#> [8648639,] 13 12 11 10 9 8 6
#> [8648640,] 13 12 11 10 9 8 7
## Factors are preserved
permuteGeneral(factor(c("low", "med", "high"),
levels = c("low", "med", "high"),
ordered = TRUE))
#> [,1] [,2] [,3]
#> [1,] low med high
#> [2,] low high med
#> [3,] med low high
#> [4,] med high low
#> [5,] high low med
#> [6,] high med low
#> Levels: low < med < highCombinations/Permutations with Repetition
There are many problems in combinatorics which require finding
combinations/permutations with repetition. This is easily achieved by
setting repetition to TRUE.
fourDays <- weekdays(as.Date("2019-10-09") + 0:3, TRUE)
ht(comboGeneral(fourDays, repetition = TRUE))
#> head -->
#> [,1] [,2] [,3] [,4]
#> [1,] "Wed" "Wed" "Wed" "Wed"
#> [2,] "Wed" "Wed" "Wed" "Thu"
#> [3,] "Wed" "Wed" "Wed" "Fri"
#> [4,] "Wed" "Wed" "Wed" "Sat"
#> [5,] "Wed" "Wed" "Thu" "Thu"
#> --------
#> tail -->
#> [,1] [,2] [,3] [,4]
#> [31,] "Fri" "Fri" "Fri" "Fri"
#> [32,] "Fri" "Fri" "Fri" "Sat"
#> [33,] "Fri" "Fri" "Sat" "Sat"
#> [34,] "Fri" "Sat" "Sat" "Sat"
#> [35,] "Sat" "Sat" "Sat" "Sat"
## When repetition = TRUE, m can exceed length(v)
ht(comboGeneral(fourDays, 8, TRUE))
#> head -->
#> [,1] [,2] [,3] [,4] [,5] [,6] [,7] [,8]
#> [1,] "Wed" "Wed" "Wed" "Wed" "Wed" "Wed" "Wed" "Wed"
#> [2,] "Wed" "Wed" "Wed" "Wed" "Wed" "Wed" "Wed" "Thu"
#> [3,] "Wed" "Wed" "Wed" "Wed" "Wed" "Wed" "Wed" "Fri"
#> [4,] "Wed" "Wed" "Wed" "Wed" "Wed" "Wed" "Wed" "Sat"
#> [5,] "Wed" "Wed" "Wed" "Wed" "Wed" "Wed" "Thu" "Thu"
#> --------
#> tail -->
#> [,1] [,2] [,3] [,4] [,5] [,6] [,7] [,8]
#> [161,] "Fri" "Fri" "Fri" "Fri" "Sat" "Sat" "Sat" "Sat"
#> [162,] "Fri" "Fri" "Fri" "Sat" "Sat" "Sat" "Sat" "Sat"
#> [163,] "Fri" "Fri" "Sat" "Sat" "Sat" "Sat" "Sat" "Sat"
#> [164,] "Fri" "Sat" "Sat" "Sat" "Sat" "Sat" "Sat" "Sat"
#> [165,] "Sat" "Sat" "Sat" "Sat" "Sat" "Sat" "Sat" "Sat"
fibonacci <- c(1L, 2L, 3L, 5L, 8L, 13L, 21L, 34L)
permsFib <- permuteGeneral(fibonacci, 5, TRUE)
ht(permsFib)
#> head -->
#> [,1] [,2] [,3] [,4] [,5]
#> [1,] 1 1 1 1 1
#> [2,] 1 1 1 1 2
#> [3,] 1 1 1 1 3
#> [4,] 1 1 1 1 5
#> [5,] 1 1 1 1 8
#> --------
#> tail -->
#> [,1] [,2] [,3] [,4] [,5]
#> [32764,] 34 34 34 34 5
#> [32765,] 34 34 34 34 8
#> [32766,] 34 34 34 34 13
#> [32767,] 34 34 34 34 21
#> [32768,] 34 34 34 34 34
## N.B. class is preserved
class(fibonacci)
#> [1] "integer"
class(permsFib[1, ])
#> [1] "integer"
## Binary representation of all numbers from 0 to 1023
ht(permuteGeneral(0:1, 10, T))
#> head -->
#> [,1] [,2] [,3] [,4] [,5] [,6] [,7] [,8] [,9] [,10]
#> [1,] 0 0 0 0 0 0 0 0 0 0
#> [2,] 0 0 0 0 0 0 0 0 0 1
#> [3,] 0 0 0 0 0 0 0 0 1 0
#> [4,] 0 0 0 0 0 0 0 0 1 1
#> [5,] 0 0 0 0 0 0 0 1 0 0
#> --------
#> tail -->
#> [,1] [,2] [,3] [,4] [,5] [,6] [,7] [,8] [,9] [,10]
#> [1020,] 1 1 1 1 1 1 1 0 1 1
#> [1021,] 1 1 1 1 1 1 1 1 0 0
#> [1022,] 1 1 1 1 1 1 1 1 0 1
#> [1023,] 1 1 1 1 1 1 1 1 1 0
#> [1024,] 1 1 1 1 1 1 1 1 1 1Working with Multisets
Sometimes, the standard combination/permutation functions don’t quite
get us to our desired goal. For example, one may need all permutations
of a vector with some of the elements repeated a specific number of
times (i.e. a multiset). Consider the following vector
a <- c(1,1,1,1,2,2,2,7,7,7,7,7) and one would like to
find permutations of a of length 6. Using traditional
methods, we would need to generate all permutations, then eliminate
duplicate values. Even considering that permuteGeneral is
very efficient, this approach is clunky and not as fast as it could be.
Observe:
getPermsWithSpecificRepetition <- function(z, n) {
b <- permuteGeneral(z, n)
myDupes <- duplicated(b)
b[!myDupes, ]
}
a <- as.integer(c(1, 1, 1, 1, 2, 2, 2, 7, 7, 7, 7, 7))
system.time(test <- getPermsWithSpecificRepetition(a, 6))
#> user system elapsed
#> 0.039 0.003 0.043Enter freqs
Situations like this call for the use of the freqs
argument. Simply, enter the number of times each unique element is
repeated and Voila!
## Using the S3 method for class 'table'
system.time(test2 <- permuteGeneral(table(a), 6))
#> user system elapsed
#> 0 0 0
identical(test, test2)
#> [1] TRUEHere are some more general examples with multisets:
## Generate all permutations of a vector with specific
## length of repetition for each element (i.e. multiset)
ht(permuteGeneral(3, freqs = c(1,2,2)))
#> head -->
#> [,1] [,2] [,3] [,4] [,5]
#> [1,] 1 2 2 3 3
#> [2,] 1 2 3 2 3
#> [3,] 1 2 3 3 2
#> [4,] 1 3 2 2 3
#> [5,] 1 3 2 3 2
#> --------
#> tail -->
#> [,1] [,2] [,3] [,4] [,5]
#> [26,] 3 2 3 1 2
#> [27,] 3 2 3 2 1
#> [28,] 3 3 1 2 2
#> [29,] 3 3 2 1 2
#> [30,] 3 3 2 2 1
## or combinations of a certain length
comboGeneral(3, 2, freqs = c(1,2,2))
#> [,1] [,2]
#> [1,] 1 2
#> [2,] 1 3
#> [3,] 2 2
#> [4,] 2 3
#> [5,] 3 3Parallel Computing
Using the parameter Parallel or nThreads,
we can generate combinations/permutations with greater efficiency.
library(microbenchmark)
## RcppAlgos uses the "number of threads available minus one" when Parallel is TRUE
RcppAlgos::stdThreadMax()
#> [1] 10
comboCount(26, 13)
#> [1] 10400600
## Compared to combn using 4 threads
microbenchmark(combn = combn(26, 13),
serAlgos = comboGeneral(26, 13),
parAlgos = comboGeneral(26, 13, nThreads = 4),
times = 10,
unit = "relative")
#> Warning in microbenchmark(combn = combn(26, 13), serAlgos = comboGeneral(26, : less
#> accurate nanosecond times to avoid potential integer overflows
#> Unit: relative
#> expr min lq mean median uq max neval
#> combn 136.390693 133.598983 98.46741 104.65877 77.056960 74.743036 10
#> serAlgos 3.168793 3.101219 2.39158 2.43135 1.788647 2.144273 10
#> parAlgos 1.000000 1.000000 1.00000 1.00000 1.000000 1.000000 10
## Using 7 cores w/ Parallel = TRUE
microbenchmark(
serial = comboGeneral(20, 10, freqs = rep(1:4, 5)),
parallel = comboGeneral(20, 10, freqs = rep(1:4, 5), Parallel = TRUE),
unit = "relative"
)
#> Unit: relative
#> expr min lq mean median uq max neval
#> serial 3.367341 3.255987 2.957451 2.718289 2.782202 2.391709 100
#> parallel 1.000000 1.000000 1.000000 1.000000 1.000000 1.000000 100Using arguments lower and upper
There are arguments lower and upper that
can be utilized to generate chunks of combinations/permutations without
having to generate all of them followed by subsetting. As the output is
in lexicographical order, these arguments specify where to start and
stop generating. For example, comboGeneral(5, 3) outputs 10
combinations of the vector 1:5 chosen 3 at a time. We can
set lower to 5 in order to start generation from the
5th lexicographical combination. Similarly, we can
set upper to 4 in order to only generate the first 4
combinations. We can also use them together to produce only a certain
chunk of combinations. For example, setting lower to 4 and
upper to 6 only produces the 4th,
5th, and 6th lexicographical
combinations. Observe:
comboGeneral(5, 3, lower = 4, upper = 6)
#> [,1] [,2] [,3]
#> [1,] 1 3 4
#> [2,] 1 3 5
#> [3,] 1 4 5
## is equivalent to the following:
comboGeneral(5, 3)[4:6, ]
#> [,1] [,2] [,3]
#> [1,] 1 3 4
#> [2,] 1 3 5
#> [3,] 1 4 5Generating Results Beyond .Machine$integer.max
In addition to being useful by avoiding the unnecessary overhead of generating all combination/permutations followed by subsetting just to see a few specific results, lower and upper can be utilized to generate large number of combinations/permutations in parallel (see this stackoverflow post for a real use case). Observe:
## Over 3 billion results
comboCount(35, 15)
#> [1] 3247943160
## 10086780 evenly divides 3247943160, otherwise you need to ensure that
## upper does not exceed the total number of results (E.g. see below, we
## would have "if ((x + foo) > 3247943160) {myUpper = 3247943160}" where
## foo is the size of the increment you choose to use in seq()).
system.time(lapply(seq(1, 3247943160, 10086780), function(x) {
temp <- comboGeneral(35, 15, lower = x, upper = x + 10086779)
## do something
x
}))
#> user system elapsed
#> 22.492 6.126 28.635
## Enter parallel
library(parallel)
system.time(mclapply(seq(1, 3247943160, 10086780), function(x) {
temp <- comboGeneral(35, 15, lower = x, upper = x + 10086779)
## do something
x
}, mc.cores = 6))
#> user system elapsed
#> 24.251 9.117 6.975GMP Support
The arguments lower and upper are also
useful when one needs to explore combinations/permutations where the
number of results is large:
set.seed(222)
myVec <- rnorm(1000)
## HUGE number of combinations
comboCount(myVec, 50, repetition = TRUE)
#> Big Integer ('bigz') :
#> [1] 109740941767310814894854141592555528130828577427079559745647393417766593803205094888320
## Let's look at one hundred thousand combinations in the range (1e15 + 1, 1e15 + 1e5)
system.time(b <- comboGeneral(myVec, 50, TRUE,
lower = 1e15 + 1,
upper = 1e15 + 1e5))
#> user system elapsed
#> 0.002 0.001 0.003
b[1:5, 45:50]
#> [,1] [,2] [,3] [,4] [,5] [,6]
#> [1,] 0.5454861 0.4787456 0.7797122 2.004614 -1.257629 -0.7740501
#> [2,] 0.5454861 0.4787456 0.7797122 2.004614 -1.257629 0.1224679
#> [3,] 0.5454861 0.4787456 0.7797122 2.004614 -1.257629 -0.2033493
#> [4,] 0.5454861 0.4787456 0.7797122 2.004614 -1.257629 1.5511027
#> [5,] 0.5454861 0.4787456 0.7797122 2.004614 -1.257629 1.0792094User Defined Functions
You can also pass user defined functions by utilizing the argument
FUN. This feature’s main purpose is for convenience,
however it is somewhat more efficient than generating all
combinations/permutations and then using a function from the
apply family (N.B. the argument Parallel has
no effect when FUN is employed).
funCustomComb = function(n, r) {
combs = comboGeneral(n, r)
lapply(1:nrow(combs), function(x) cumprod(combs[x,]))
}
identical(funCustomComb(15, 8), comboGeneral(15, 8, FUN = cumprod))
#> [1] TRUE
microbenchmark(f1 = funCustomComb(15, 8),
f2 = comboGeneral(15, 8, FUN = cumprod), unit = "relative")
#> Unit: relative
#> expr min lq mean median uq max neval
#> f1 5.235565 5.189387 5.615276 5.172509 5.101059 20.59012 100
#> f2 1.000000 1.000000 1.000000 1.000000 1.000000 1.00000 100
comboGeneral(15, 8, FUN = cumprod, upper = 3)
#> [[1]]
#> [1] 1 2 6 24 120 720 5040 40320
#>
#> [[2]]
#> [1] 1 2 6 24 120 720 5040 45360
#>
#> [[3]]
#> [1] 1 2 6 24 120 720 5040 50400
## An example involving the powerset... Note, we could
## have used the FUN.VALUE parameter here instead of
## calling unlist. See the next section.
unlist(comboGeneral(c("", letters[1:3]), 3,
freqs = c(2, rep(1, 3)),
FUN = function(x) paste(x, collapse = "")))
#> [1] "a" "b" "c" "ab" "ac" "bc" "abc"Using FUN.VALUE
As of version 2.5.0, we can make use of
FUN.VALUE which serves as a template for the return value
from FUN. The behavior is nearly identical to
vapply:
## Example from earlier involving the power set
comboGeneral(c("", letters[1:3]), 3, freqs = c(2, rep(1, 3)),
FUN = function(x) paste(x, collapse = ""), FUN.VALUE = "a")
#> [1] "a" "b" "c" "ab" "ac" "bc" "abc"
comboGeneral(15, 8, FUN = cumprod, upper = 3, FUN.VALUE = as.numeric(1:8))
#> [,1] [,2] [,3] [,4] [,5] [,6] [,7] [,8]
#> [1,] 1 2 6 24 120 720 5040 40320
#> [2,] 1 2 6 24 120 720 5040 45360
#> [3,] 1 2 6 24 120 720 5040 50400
## Fun example with binary representations... consider the following:
permuteGeneral(0:1, 3, TRUE)
#> [,1] [,2] [,3]
#> [1,] 0 0 0
#> [2,] 0 0 1
#> [3,] 0 1 0
#> [4,] 0 1 1
#> [5,] 1 0 0
#> [6,] 1 0 1
#> [7,] 1 1 0
#> [8,] 1 1 1
permuteGeneral(c(FALSE, TRUE), 3, TRUE, FUN.VALUE = 1,
FUN = function(x) sum(2^(which(rev(x)) - 1)))
#> [1] 0 1 2 3 4 5 6 7Passing additional arguments with ...
As of version 2.8.3, we have added the ability to pass
further arguments to FUN via ....
## Again, same example with the power set only this time we
## conveniently pass the additional arguments to paste via '...'
comboGeneral(c("", letters[1:3]), 3, freqs = c(2, rep(1, 3)),
FUN = paste, collapse = "", FUN.VALUE = "a")
#> [1] "a" "b" "c" "ab" "ac" "bc" "abc"This concludes our discussion around user defined functions. There are several nice features that allow the user to more easily get the desired output with fewer function calls as well as fewer keystrokes. This was most clearly seen in our example above with the power set.
We started with wrapping our call to comboGeneral with
unlist, which was alleviated by the parameter
FUN.VALUE. We then further simplified our usage of
FUN by allowing additional arguments to be passed via
....
S3 methods
As of version 2.8.3, we have added several S3 methods
for convenience.
Take our earlier example where we were talking about multisets.
a <- as.integer(c(1, 1, 1, 1, 2, 2, 2, 7, 7, 7, 7, 7))
## Explicitly utilizing the freqs argument and determining the unique
## values for v... Still works, but clunky
t1 <- permuteGeneral(rle(a)$values, 6, freqs = rle(a)$lengths)
## Now using the table method... much cleaner
t2 <- permuteGeneral(table(a), 6)
identical(t1, t2)
#> [1] TRUEThere are other S3 methods defined that simplify the interface. Take
for example the case when we want to pass a character vector. We know
underneath the hood, character vectors are not thread safe so the
Parallel and nThreads argument are ignored. We
also know that the constraints parameters are only applicable to numeric
vectors. For these reason, our default method’s interface is greatly
simplified:
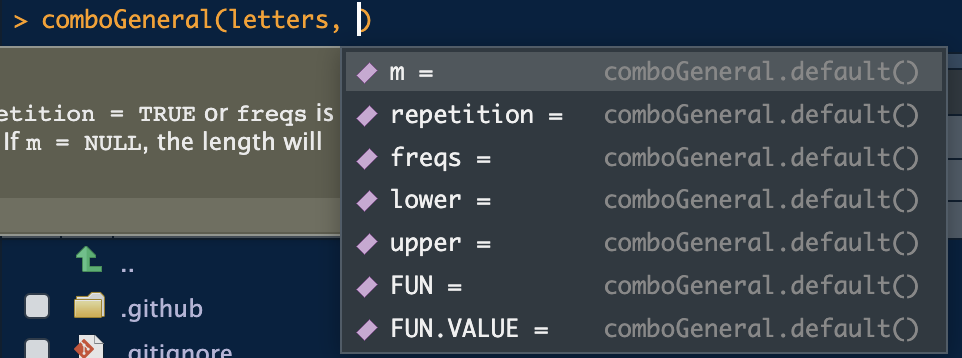
We see only the necessary options. With numeric types, the options are more numerous:
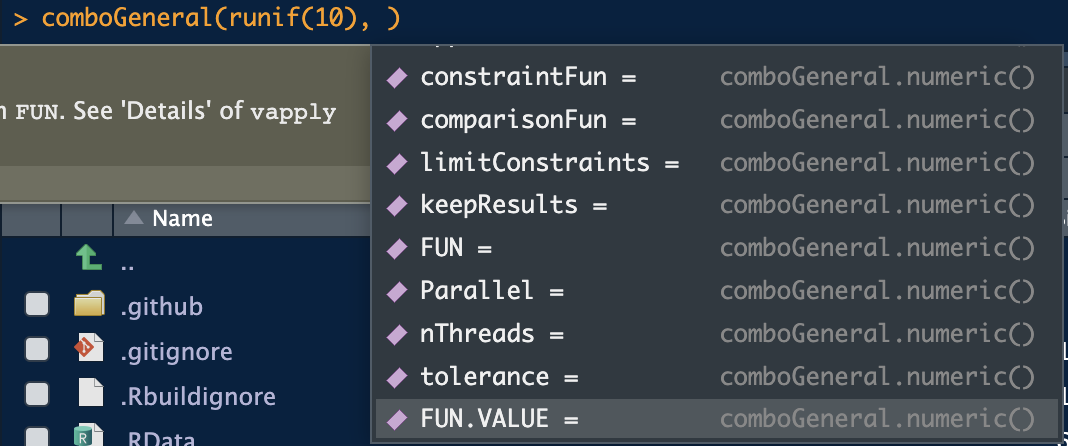
There is also a list method that allows one to find
combinations or permutations of lists:
comboGeneral(
list(
numbers = rnorm(4),
states = state.abb[1:5],
some_data = data.frame(a = c('a', 'b'), b = c(10, 100))
),
m = 2
)
#> [[1]]
#> [[1]]$numbers
#> [1] -1.9376332 0.2583997 -0.7198657 -0.8985872
#>
#> [[1]]$states
#> [1] "AL" "AK" "AZ" "AR" "CA"
#>
#>
#> [[2]]
#> [[2]]$numbers
#> [1] -1.9376332 0.2583997 -0.7198657 -0.8985872
#>
#> [[2]]$some_data
#> a b
#> 1 a 10
#> 2 b 100
#>
#>
#> [[3]]
#> [[3]]$states
#> [1] "AL" "AK" "AZ" "AR" "CA"
#>
#> [[3]]$some_data
#> a b
#> 1 a 10
#> 2 b 100This feature was inspired by ggrothendieck here: Issue 20.